TEV protease
| nuclear-inclusion-a endopeptidase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
|
| |||||||||
| Identifiers | |||||||||
| EC number | 3.4.22.44 | ||||||||
| CAS number | 139946-51-3 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| |||||||||
TEV protease (EC 3.4.22.44, Tobacco Etch Virus nuclear-inclusion-a endopeptidase) is a highly sequence-specific cysteine protease from Tobacco Etch Virus (TEV).[1] It is a member of the PA clan of chymotrypsin-like proteases.[2] Due to its high sequence specificity it is frequently used for the controlled cleavage of fusion proteins in vitro and in vivo.[3]
Origin
The tobacco etch virus encodes its entire genome as a single massive polyprotein (350 kDa). This is cleaved into functional units by the three proteases: P1 protease (1 cleavage site), helper-component protease (1 cleavage site) and TEV protease (7 cleavage sites).[1] The native protease also contains an internal self-cleavage site. This site is slowly cleaved to inactivate the enzyme (the physiological reason for this is unknown).
Structure and Function
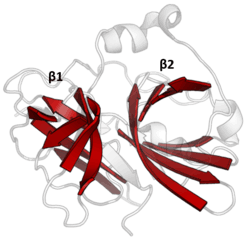
The structure of TEV protease has been solved by X-ray crystallography.[4] It is composed of two β-barrels and a flexible C-terminal tail and displays structural homology to the chymotrypsin superfamily of proteases (PA clan, C4 family by MEROPS classification).[2] Although homologous to cellular serine proteases (such as trypsin, elastase, thrombin etc.), TEV protease uses a cysteine as its catalytic nucleophile[5] (as do many other viral proteases).
Covalent catalysis is performed with an Asp-His-Cys triad, split between the two barrels (Asp on β1 and His and Cys on β2).[6] The substrate is held as a β-sheet, forming an antiparallel interaction with the cleft between the barrels and a parallel interaction with the C-terminal tail.[7] The enzyme therefore forms a binding tunnel around the substrate and side chain interactions control specificity.[4]
Specificity

The preferred, native cleavage sequence was first identified by examining the cut sites in the native polyprotein substrate for recurring sequence. The consensus for these native cut sites is ENLYFQ\S where ‘\’ denotes the cleaved peptide bond.[8] Residues of the substrate are labelled P6 to P1 before the cut site and P1’ after the cut site. Early works also measured cleavage of an array of similar substrates to characterise how specific the protease was for the native sequence.[9][10]
Studies have subsequently used sequencing of cleaved substrates from a pool of randomised sequences to determine preference patterns.[11][12] Although ENLYFQ\S is the optimal sequence, the protease is active to a greater or lesser extent on a range of substrates (i.e. shows some substrate promiscuity). The highest cleavage is of sequences closest to the consensus EXLYΦQ\φ where X is any residue, Φ is any large or medium hydrophobe and φ is any small hydrophobic or polar residue.
Specificity is endowed by the large contact area between enzyme and substrate. Proteases such as trypsin have specificity for one residue before and after the cleaved bond due to a shallow binding cleft with only one or two pockets that bind the substrate side chains. Conversely, viral proteases such as TEV protease have a long C-terminal tail which completely covers the substrate to create a binding tunnel. This tunnel contains a set of tight binding pockets such that each side chain of the substrate peptide (P6 to P1’) is bound in a complementary site (S6 to S1’).[4]
In particular, peptide side chain P6-Glu contacts a network of three hydrogen bonds; P5-Asn points into the solvent, making no specific interactions (hence the absence of substrate consensus at this position); P4-Leu is buried in a hydrophobic pocket; P3-Tyr is held in a hydrophobic pocket with a short hydrogen bond at the end; P2-Phe is also surrounded by hydrophobes including the face of the triad histidine; P1-Gln forms four hydrogen bonds; and P1’-Ser is only partly enclosed in a shallow hydrophobic groove.[4]
As a biochemical tool
One of the main uses of this protein is for removing affinity tags from purified recombinant fusion proteins. The reason for the use of TEV protease as a biochemical tool is its high sequence specificity. This specificity allows for the controlled cleavage of proteins when the preference sequence is inserted into flexible loops. It also makes it relatively non-toxic in vivo as the recognized sequence scarcely occurs in proteins.[13]
Although rational design has had limited success in changing protease specificity, directed evolution has been used to change the preferred residue either before[14] or after[15][16] the cleavage site.
However, TEV protease does have limitations as a biochemical tool. It is prone to deactivation by self-cleavage (autolysis), though this can be abolished through a single S219V mutation in the internal cleavage site.[17] The protease expressed alone is also poorly soluble, however several attempts have been made to improve its solubility through directed evolution and computational design. It has also been shown that expression can be improved by fusion to Maltose Binding Protein (MBP) which acts a solubility enhancing partner.
The molecular weight of this enzyme varies between 25 and 27 kDa depending on the specific construct used.
External links
References
- 1 2 UniProt: TEV polyprotein: "P04517".
- 1 2 Rawlings ND, Barrett AJ, Bateman A (January 2012). "MEROPS: the database of proteolytic enzymes, their substrates and inhibitors". Nucleic Acids Res. 40 (Database issue): D343–50. doi:10.1093/nar/gkr987. PMC 3245014
 . PMID 22086950.
. PMID 22086950. - ↑ Kapust RB, Waugh DS (July 2000). "Controlled intracellular processing of fusion proteins by TEV protease". Protein Expr. Purif. 19 (2): 312–8. doi:10.1006/prep.2000.1251. PMID 10873547.
- 1 2 3 4 Phan J, Zdanov A, Evdokimov AG, Tropea JE, Peters HK, Kapust RB, Li M, Wlodawer A, Waugh DS (December 2002). "Structural basis for the substrate specificity of tobacco etch virus protease". J. Biol. Chem. 277 (52): 50564–72. doi:10.1074/jbc.M207224200. PMID 12377789.
- ↑ Bazan JF, Fletterick RJ (November 1988). "Viral cysteine proteases are homologous to the trypsin-like family of serine proteases: structural and functional implications". Proc. Natl. Acad. Sci. U.S.A. 85 (21): 7872–6. Bibcode:1988PNAS...85.7872B. doi:10.1073/pnas.85.21.7872. PMC 282299
 . PMID 3186696.
. PMID 3186696. - ↑ Dougherty WG, Parks TD, Cary SM, Bazan JF, Fletterick RJ (September 1989). "Characterization of the catalytic residues of the tobacco etch virus 49-kDa proteinase". Virology. 172 (1): 302–10. doi:10.1016/0042-6822(89)90132-3. PMID 2475971.
- ↑ Tyndall JD, Nall T, Fairlie DP (March 2005). "Proteases universally recognize beta strands in their active sites". Chem. Rev. 105 (3): 973–99. doi:10.1021/cr040669e. PMID 15755082.
- ↑ Carrington JC, Dougherty WG (May 1988). "A viral cleavage site cassette: identification of amino acid sequences required for tobacco etch virus polyprotein processing". Proc. Natl. Acad. Sci. U.S.A. 85 (10): 3391–5. Bibcode:1988PNAS...85.3391C. doi:10.1073/pnas.85.10.3391. PMC 280215
 . PMID 3285343.
. PMID 3285343. - ↑ Dougherty WG, Cary SM, Parks TD (August 1989). "Molecular genetic analysis of a plant virus polyprotein cleavage site: a model". Virology. 171 (2): 356–64. doi:10.1016/0042-6822(89)90603-X. PMID 2669323.
- ↑ Kapust RB, Tözsér J, Copeland TD, Waugh DS (June 2002). "The P1' specificity of tobacco etch virus protease". Biochem. Biophys. Res. Commun. 294 (5): 949–55. doi:10.1016/S0006-291X(02)00574-0. PMID 12074568.
- ↑ Boulware KT, Jabaiah A, Daugherty PS (June 2010). "Evolutionary optimization of peptide substrates for proteases that exhibit rapid hydrolysis kinetics". Biotechnol. Bioeng. 106 (3): 339–46. doi:10.1002/bit.22693. PMID 20148412.
- ↑ Kostallas G, Löfdahl P, Samuelson P (2011). "Substrate profiling of tobacco etch virus protease using a novel fluorescence-assisted whole-cell assay". PLoS ONE. 6 (1): e16136. doi:10.1371/journal.pone.0016136. PMC 3022733
 . PMID 21267463.
. PMID 21267463. - ↑ Parks TD, Leuther KK, Howard ED, Johnston SA, Dougherty WG (February 1994). "Release of proteins and peptides from fusion proteins using a recombinant plant virus proteinase". Anal. Biochem. 216 (2): 413–7. doi:10.1006/abio.1994.1060. PMID 8179197.
- ↑ Yi L, Gebhard MC, Li Q, Taft JM, Georgiou G, Iverson BL (April 2013). "Engineering of TEV protease variants by yeast ER sequestration screening (YESS) of combinatorial libraries". Proc. Natl. Acad. Sci. U.S.A. 110 (18): 7229–34. doi:10.1073/pnas.1215994110. PMC 3645551
 . PMID 23589865.
. PMID 23589865. - ↑ Renicke C, Spadaccini R, Taxis C (2013). "A tobacco etch virus protease with increased substrate tolerance at the P1' position". PLoS ONE. 8 (6): e67915. doi:10.1371/journal.pone.0067915. PMC 3691164
 . PMID 23826349.
. PMID 23826349. - ↑ Verhoeven KD, Altstadt OC, Savinov SN (March 2012). "Intracellular detection and evolution of site-specific proteases using a genetic selection system". Appl. Biochem. Biotechnol. 166 (5): 1340–54. doi:10.1007/s12010-011-9522-6. PMID 22270548.
- ↑ Kapust RB, Tözsér J, Fox JD, Anderson DE, Cherry S, Copeland TD, Waugh DS (December 2001). "Tobacco etch virus protease: mechanism of autolysis and rational design of stable mutants with wild-type catalytic proficiency". Protein Eng. 14 (12): 993–1000. doi:10.1093/protein/14.12.993. PMID 11809930.