Spliceosome
A spliceosome is a large and complex molecular machine found primarily within the splicing speckles of the cell nucleus of eukaryotic cells. The spliceosome is assembled from snRNAs and protein complexes. The spliceosome removes introns from a transcribed pre-mRNA, a type of primary transcript. This process is generally referred to as splicing.[1] Only eukaryotes have spliceosomes and some organisms have a second spliceosome, the minor spliceosome.[2] An analogy is a film editor, who selectively cuts out irrelevant or incorrect material (equivalent to the introns) from the dailies and sends the cleaned-up version to be screened for the producer.

Composition
Each spliceosome is composed of five small nuclear RNAs (snRNA), and a range of associated protein factors. When these small RNA are combined with the protein factors, they make an RNA-protein complex called snRNP (small nuclear ribonucleo proteins). The snRNAs that make up the major spliceosome are named U1, U2, U4, U5, and U6, and participate in several RNA-RNA and RNA-protein interactions. The RNA component of the small nuclear ribonucleic protein or snRNP (pronounced "snurp") is rich in uridine (the nucleoside analog of the uracil nucleotide).
The canonical assembly of the spliceosome occurs anew on each hnRNA (pre-mRNA, heterogeneous nuclear RNA). The hnRNA contains specific sequence elements that are recognized and utilized during spliceosome assembly. These include the 5' end splice, the branch point sequence, the polypyrimidine tract, and the 3' end splice site. The spliceosome catalyzes the removal of introns, and the ligation of the flanking exons.
Introns typically have a GU nucleotide sequence at the 5' end splice site, and an AG at the 3' end splice site. The 3' splice site can be further defined by a variable length of polypyrimidines, called the polypyrimidine tract (PPT), which serves the dual function of recruiting factors to the 3' splice site and possibly recruiting factors to the branch point sequence (BPS). The BPS contains the conserved Adenosine required for the first step of splicing.
A group of less abundant snRNAs, U11, U12, U4atac, and U6atac, together with U5, are subunits of the so-called minor spliceosome that splices a rare class of pre-mRNA introns, denoted U12-type. The minor spliceosome is located in the nucleus like its major counterpart,[3] though there are exceptions in some specialised cells including anucleate platelets[4] and the dendroplasm (dendrite cytoplasm) of neuronal cells.[5]
New evidence derived from the first crystal structure of a group II intron suggests that the spliceosome is actually a ribozyme, and that it uses a two–metal ion mechanism for catalysis.[6] In addition, many proteins exhibit a zinc-binding motif, which underscores the importance of zinc metal in the splicing mechanism.[7][8][9] The first molecular-resolution reconstruction of U4/U6.U5 triple small nuclear ribonucleoprotein (tri-snRNP) complex was reported in 2016.[10]
Alternative splicing
Alternative splicing (the re-combination of different exons) is a major source of genetic diversity in eukaryotes. Splice variants have been used to account for the relatively small number of genes in the human genome. For years the estimate widely varied, with top estimates reaching 100,000 genes,[12] but now, due to the Human Genome Project, the figure is believed to be closer to 20,000 genes. One particular Drosophila gene (Dscam, the Drosophila homolog of the human Down syndrome cell adhesion molecule DSCAM) can be alternatively spliced into 38,000 different mRNA.[13]
RNA splicing
In 1977, work by the Sharp and Roberts labs revealed that genes of higher organisms are "split" or present in several distinct segments along the DNA molecule.[14][15] The coding regions of the gene are separated by non-coding DNA that is not involved in protein expression. The split gene structure was found when adenoviral mRNAs were hybridized to endonuclease cleavage fragments of single stranded viral DNA.[14] It was observed that the mRNAs of the mRNA-DNA hybrids contained 5' and 3' tails of non-hydrogen bonded regions. When larger fragments of viral DNAs were used, forked structures of looped out DNA were observed when hybridized to the viral mRNAs. It was realized that the looped out regions, the introns, are excised from the precursor mRNAs in a process Sharp named "splicing". The split gene structure was subsequently found to be common to most eukaryotic genes. Phillip Sharp and Richard J. Roberts were awarded the 1993 Nobel Prize in Physiology or Medicine for their discovery of introns and the splicing process.
Spliceosome assembly
The model for formation of the spliceosome active site involves an ordered, stepwise assembly of discrete snRNP particles on the hnRNA substrate. The first recognition of hnRNAs involves U1 snRNP binding to the 5' end splice site of the hnRNA and other non-snRNP associated factors to form the commitment complex, or early (E) complex in mammals.[16][17] The commitment complex is an ATP-independent complex that commits the hnRNA to the splicing pathway.[18] U2 snRNP is recruited to the branch region through interactions with the E complex component U2AF (U2 snRNP auxiliary factor) and possibly U1 snRNP. In an ATP-dependent reaction, U2 snRNP becomes tightly associated with the branch point sequence (BPS) to form complex A. A duplex formed between U2 snRNP and the hnRNA branch region bulges out the branch adenosine specifying it as the nucleophile for the first transesterification.[19]
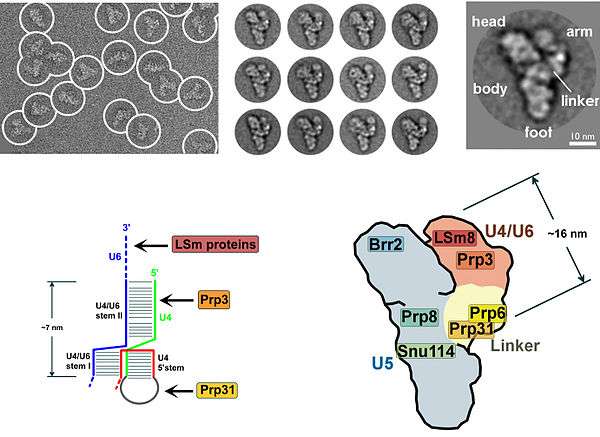
The presence of a pseudouridine residue in U2 snRNA, nearly opposite of the branch site, results in an altered conformation of the RNA-RNA duplex upon the U2 snRNP binding. Specifically, the altered structure of the duplex induced by the pseudouridine places the 2' OH of the bulged adenosine in a favorable position for the first step of splicing.[20] The U4/U5/U6 tri-snRNP (see Figure 1) is recruited to the assembling spliceosome to form complex B, and following several rearrangements, complex C (the spliceosome) is activated for catalysis.[21][22] It is unclear how the triple snRNP is recruited to complex A, but this process may be mediated through protein-protein interactions and/or base pairing interactions between U2 snRNA and U6 snRNA.
The U5 snRNP interacts with sequences at the 5' and 3' splice sites via the invariant loop of U5 snRNA[23] and U5 protein components interact with the 3' splice site region.[24]
Upon recruitment of the triple snRNP, several RNA-RNA rearrangements precede the first catalytic step and further rearrangements occur in the catalytically active spliceosome. Several of the RNA-RNA interactions are mutually exclusive; however, it is not known what triggers these interactions, nor the order of these rearrangements. The first rearrangement is probably the displacement of U1 snRNP from the 5' splice site and formation of a U6 snRNA interaction. It is known that U1 snRNP is only weakly associated with fully formed spliceosomes,[25] and U1 snRNP is inhibitory to the formation of a U6-5' splice site interaction on a model of substrate oligonucleotide containing a short 5' exon and 5' splice site.[26] Binding of U2 snRNP to the branch point sequence (BPS) is one example of an RNA-RNA interaction displacing a protein-RNA interaction. Upon recruitment of U2 snRNP, the branch binding protein SF1 in the commitment complex is displaced since the binding site of U2 snRNA and SF1 are mutually exclusive events.
Within the U2 snRNA, there are other mutually exclusive rearrangements that occur between competing conformations. For example, in the active form, stem loop IIa is favored; in the inactive form a mutually exclusive interaction between the loop and a downstream sequence predominates.[22] It is unclear how U4 is displaced from U6 snRNAm, although RNA has been implicated in spliceosome assembly, and may function to unwind U4/U6 and promote the formation of a U2/U6 snRNA interaction. The interactions of U4/U6 stem loops I and II dissociate and the freed stem loop II region of U6 folds on itself to form an intramolecular stem loop and U4 is no longer required in further spliceosome assembly. The freed stem loop I region of U6 base pairs with U2 snRNA forming the U2/U6 helix I. However, the helix I structure is mutually exclusive with the 3' half of an internal 5' stem loop region of U2 snRNA.
References
- ↑ Will, Cindy L.; Reinhard Lührmann (2011-07-01). "Spliceosome Structure and Function". Cold Spring Harbor Perspectives in Biology. 3 (7): a003707. doi:10.1101/cshperspect.a003707. Retrieved 2012-01-25.
- ↑ Patel, Abhijit A.; Joan A. Steitz (December 2003). "Splicing double: insights from the second spliceosome". Nature Reviews Molecular Cell Biology. 4 (12): 960–970. doi:10.1038/nrm1259. ISSN 1471-0072. PMID 14685174. Retrieved 2013-07-24.
- ↑ Pessa, HK; Will, CL; Meng, X; Schneider, C; Watkins, NJ; Perälä, N; Nymark, M; Turunen, JJ; Lührmann, R; Frilander, MJ (Jun 24, 2008). "Minor spliceosome components are predominantly localized in the nucleus.". Proceedings of the National Academy of Sciences of the United States of America. 105 (25): 8655–60. doi:10.1073/pnas.0803646105. PMC 2438382
 . PMID 18559850.
. PMID 18559850. - ↑ Denis, MM; Tolley, ND; Bunting, M; Schwertz, H; Jiang, H; Lindemann, S; Yost, CC; Rubner, FJ; Albertine, KH; Swoboda, KJ; Fratto, CM; Tolley, E; Kraiss, LW; McIntyre, TM; Zimmerman, GA; Weyrich, AS (Aug 12, 2005). "Escaping the nuclear confines: signal-dependent pre-mRNA splicing in anucleate platelets.". Cell. 122 (3): 379–91. doi:10.1016/j.cell.2005.06.015. PMID 16096058. Cite uses deprecated parameter
|coauthors=(help) - ↑ Glanzer, J; Miyashiro, KY; Sul, JY; Barrett, L; Belt, B; Haydon, P; Eberwine, J (Nov 15, 2005). "RNA splicing capability of live neuronal dendrites.". Proceedings of the National Academy of Sciences of the United States of America. 102 (46): 16859–64. doi:10.1073/pnas.0503783102. PMC 1277967
 . PMID 16275927.
. PMID 16275927. - ↑ Toor N, Keating KS, Taylor SD, Pyle AM (2008). "Crystal structure of a self-spliced group II intron". Science. 320 (5872): 77–82. doi:10.1126/science.1153803. PMID 18388288.
- ↑ Agafonov, DE; Deckert, J; Wolf, E; Odenwa'Ider, P; Bessonov, S; Will, CL; Urlaub, H; Lu'hrmann, R (2011). "Semiquantitative proteomic analysis of the human spliceosome via a novel two-dimensional gel electrophoresis method". Mol Cell Biol. 31: 2667–2682. doi:10.1128/mcb.05266-11.
- ↑ Kuhn, AN; van Santen, MA; Schwienhorst, A; Urlaub, H; Lu'hrmann, R (2009). "Stalling of spliceosome assembly at distinct stages by small molecule inhibitors of protein acetylation and deacetylation". RNA. 15: 153–175. doi:10.1261/rna.1332609.
- ↑ Patil, V.; Canzoneri, J.; Samatov, T.; Luhrmann, R.; Oyelere, A. K. (2012). "Molecular architecture of zinc chelating small molecules that stall spliceosome assembly at distinct stages". RNA. 18 (9): 1605–1611. doi:10.1261/rna.034819.112.
- ↑ Cate, Jamie H. D. (2016-03-25). "A Big Bang in spliceosome structural biology". Science. 351 (6280): 1390–1392. doi:10.1126/science.aaf4465. ISSN 0036-8075. PMID 27013712.
- ↑ Häcker I, Sander B, Golas MM, Wolf E, Karagöz E, Kastner B, Stark H, Fabrizio P, Lührmann R (2008). "Localization of Prp8, Brr2, Snu114 and U4/U6 proteins in the yeast tri-snRNP by electron microscopy". Nat Struct Mol Biol. 15 (11): 1206–12. doi:10.1038/nsmb.1506. PMID 18953335.
- ↑ Smaglik, P. (2000). "Researchers take a gamble on the human genome". Nature. 405 (6784): 264. doi:10.1038/35012771. PMID 10830930.
- ↑ Schmucker, D.; Clemens, J.C.; Shu, H.; Worby, C.A.; Xiao, J.; Muda, M.; Dixon, J.E.; Zipursky, S.L. (2000). "Drosophila Dscam Is an Axon Guidance Receptor Exhibiting Extraordinary Molecular Diversity". Cell. 101 (6): 671–684. doi:10.1016/S0092-8674(00)80878-8. PMID 10892653.
- 1 2 Berget, S. M., Moore, C. and Sharp, P. A. (1977). "Spliced segments at the 5' terminus of adenovirus 2 late mRNA". Proc. Natl. Acad. Sci. USA. 74 (8): 3171–5. doi:10.1073/pnas.74.8.3171. PMC 431482
 . PMID 269380.
. PMID 269380. - ↑ Chow, L. T.; Roberts, J. M.; Lewis, J. B.; Broker, T. R. (1977). "A map of cytoplasmic RNA transcripts from lytic adenovirus type 2, determined by electron microscopy of RNA:DNA hybrids". Cell. 11 (4): 819–836. doi:10.1016/0092-8674(77)90294-X. PMID 890740.
- ↑ Jamison SF, Crow A, Garcia-Blanco MA (October 1, 1992). "The spliceosome assembly pathway in mammalian extracts". Molecular and Cell Biology. 12 (10): 4279–87. PMC 360351
 . PMID 1383687.
. PMID 1383687. - ↑ Seraphin B. & Rosbash M. (1989). "Identification of functional U1 snRNA pre-messenger RNA complexes committed to spliceosome assembly and splicing". Cell. 59 (2): 349–58. doi:10.1016/0092-8674(89)90296-1. PMID 2529976.
- ↑ Legrain P, Seraphin B, Rosbash M (September 1, 1988). "Early commitment of yeast pre-mRNA to the spliceosome pathway". Mol. Cell. Biol. 8 (9): 3755–60. PMC 365433
 . PMID 3065622.
. PMID 3065622. - ↑ Query, C. C., M. J. Moore, and P. Sharp (1994). "Branch nucleophile selection in pre-mRNA splicing: evidence for the bulged duplex model". Genes Dev. 8 (5): 587–97. doi:10.1101/gad.8.5.587. PMID 7926752.
- ↑ Newby M. I. & Greenbaum, N. L. (2002). "Sculpting of the spliceosomal branch site recognition motif by a conserved pseudouridine". Nature Structural Biology. 9 (12): 958–65. doi:10.1038/nsb873. PMID 12426583.
- ↑ Burge, C.B.; et al. (1999). "Splicing precursors to mRNAs by the spliceosomes". In Gesteland, R.F.; Cech, T.R.; Atkins, J.F. The RNA World. Cold Spring Harbor Lab. Press. pp. 525–60. ISBN 0-87969-380-0.
- 1 2 Staley JP, Guthrie C (1998). "Mechanical devices of the spliceosome: motors, clocks, springs, and things". Cell. 92 (3): 315–26. doi:10.1016/S0092-8674(00)80925-3. PMID 9476892.
- ↑ Newman AJ, Teigelkamp S, Beggs JD (1995). "snRNA interactions at 5' and 3' splice sites monitored by photoactivated crosslinking in yeast spliceosomes". RNA. 1 (9): 968–80. PMC 1369345
 . PMID 8548661.
. PMID 8548661. - ↑ Chiara MD, Palandjian L, Feld Kramer R, Reed R (1997). "Evidence that U5 snRNP recognizes the 3' splice site for catalytic step II in mammals". EMBO J. 16 (15): 4746–59. doi:10.1093/emboj/16.15.4746. PMC 1170101
 . PMID 9303319.
. PMID 9303319. - ↑ Moore, M. J. & Sharp, P. A. (1993). "Evidence for two active sites in the spliceosome provided by stereochemistry of pre-mRNA splicing". Nature. 365 (6444): 364–8. doi:10.1038/365364a0. PMID 8397340.
- ↑ Konforti BB, Koziolkiewicz MJ, Konarska MM (1993). "Disruption of base pairing between the 5' splice site and the 5' end of U1 snRNA is required for spliceosome assembly". Cell. 75 (5): 863–73. doi:10.1016/0092-8674(93)90531-T. PMID 8252623.
External links
| Wikimedia Commons has media related to Spliceosomes. |
- Butcher, Samuel E. (2011). "Chapter 8. The Spliceosome and Its Metal Ions". In Astrid Sigel; Helmut Sigel; Roland K. O. Sigel. Structural and catalytic roles of metal ions in RNA. Metal Ions in Life Sciences. 9. RSC Publishing. pp. 235–51. doi:10.1039/9781849732512-00235.
- Nilsen T (2003). "The spliceosome: the most complex macromolecular machine in the cell?". BioEssays. 25 (12): 1147–9. doi:10.1002/bies.10394. PMID 14635248.
- Spliceosomes at the US National Library of Medicine Medical Subject Headings (MeSH)
- 3D macromolecular structures of Spliceosomes from the EM Data Bank(EMDB)