Inner cell mass
| Inner cell mass | |
|---|---|
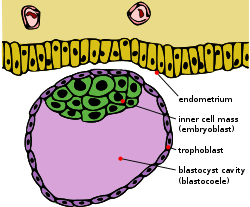 Blastocyst with an inner cell mass and trophoblast. | |
| Details | |
| Carnegie stage | 3 |
| Days | 6 |
| Precursor | blastocyst |
| Gives rise to | epiblast, hypoblast |
| Identifiers | |
| Latin | embryoblastus; massa cellularis interna; pluriblastus senior |
| TE | E6.0.1.1.2.0.4 |
| FMA | 86557 |
In early embryogenesis of most eutherian mammals, the inner cell mass (abbreviated ICM and also known as the embryoblast in mammals or pluriblast) is the mass of cells inside the primordial embryo that, once fertilized, will eventually give rise to the definitive structures of the fetus. This structure forms in the earliest steps of development, before implantation into the endometrium of the uterus has occurred. The ICM lies within the blastocoele (more correctly termed "blastocyst cavity," as it is not strictly homologous to the blastocoele of anamniote vertebrates) and is entirely surrounded by the single layer of cells called trophoblast.
Further development
The physical and functional separation of the inner cell mass from the trophectoderm (TE) is a special feature of mammalian development and is the first cell lineage specification in these embryos. Following fertilization in the oviduct, the mammalian embryo undergoes a relatively slow round of cleavages to produce an eight cell morula. Each cell of the morula, called a blastomere, increases surface contact with its neighbors in a process called compaction. This results in a polarization of the cells within the morula, and further cleavage yields a blastocyst of roughly 32 cells.[1] In mice, about 12 internal cells comprise the new inner cell mass and 20 – 24 cells comprise the surrounding trophectoderm.[2][3] There is variation between species of mammals as to number of cells at compaction with bovine embryos showing differences related to compaction as early as 9-15 cells and in rabbits not until after 32 cells.[4] There is also interspecies variation in gene expression patterns in early embryos [5]
The ICM and the TE will generate distinctly different cell types as implantation starts and embryogenesis continues. Trophectoderm cells form extraembryonic tissues, which act in a supporting role for the embryo proper. Furthermore, these cells pump fluid into the interior of the blastocyst, causing the formation of a polarized blastocyst with the ICM attached to the trophectoderm at one end (see figure). This difference in cellular localization causes the ICM cells exposed to the fluid cavity to adopt a primitive endoderm (or hypoblast) fate, while the remaining cells adopt a primitive ectoderm (or epiblast) fate. The hypoblast contributes to extraembryonic membranes and the epiblast will give rise to the ultimate embryo proper as well as some extraembryonic tissues.[1]
Regulation of cellular specification
Since segregation of pluripotent cells of the inner cell mass from the remainder of the blastocyst is integral to mammalian development, considerable research has been performed to elucidate the corresponding cellular and molecular mechanisms of this process. There is primary interest in which transcription factors and signaling molecules direct blastomere asymmetric divisions leading to what are known as inside and outside cells and thus cell lineage specification. However, due to the variability and regulative nature of mammalian embryos, experimental evidence for establishing these early fates remains incomplete.[2]
At the transcription level, the transcription factors Oct4, Nanog, Cdx2, and Tead4 have all been implicated in establishing and reinforcing the specification of the ICM and the TE in early mouse embryos.[2]

- Oct4: Oct4 is expressed in the ICM and participate in maintaining its pluripotency, a role that has been recapitulated in ICM derived mouse embryonic stem cells.[6] Oct4 genetic knockout cells both in vivo and in culture display TE morphological characteristics. It has been shown that one transcriptional target of Oct4 is the Fgf4 gene. This gene normally encodes a ligand secreted by the ICM, which induces proliferation in the adjacent polar TE.[6]
- Nanog: Nanog is also expressed in the ICM and participates in maintaining its pluripotency. In contrast with Oct4, studies of Nanog-null mice do not show the reversion of the ICM to a TE-like morphology, but demonstrate that loss of Nanog prevents the ICM from generating primitive endoderm.[7]
- Cdx2: Cdx2 is strongly expressed in the TE and is required for maintaining its specification. Knockout mice for the Cdx2 gene undergo compaction, but lose the TE epithelial integrity during the late blastocyst stage. Furthermore, Oct4 expression is subsequently raised in these TE cells, indicating Cdx2 plays a role in suppressing Oct4 in this cell lineage. Moreover, embryonic stem cells can be generated from Cdx2-null mice, demonstrating that Cdx2 is not essential for ICM specification.[8]
- Tead4: Like Cdx2, Tead4 is required for TE function, although the transcription factor is expressed ubiquitously. Tead4-null mice similarly undergo compaction, but fail to generate the blastocoel cavity. Like Cdx2-null embryos, the Tead4-null embryos can yield embryonic stem cells, indicating that Tead4 is dispensable for ICM specification.[9] Recent work has shown that Tead4 may help to upregulate Cdx2 in the TE and its transcriptional activity depends on the coactivator Yap. Yap’s nuclear localization in outside cells allows it to contribute to TE specificity, whereas inside cells sequester Yap in the cytoplasm through a phosphorylation event.[10]
Together these transcription factors function in a positive feedback loop that strengthens the ICM to TE cellular allocation. Initial polarization of blastomeres occurs at the 8-16 cell stage. An apical-basolateral polarity is visible through the visualization of apical markers such as Par3, Par6, and aPKC as well as the basal marker E-Cadherin.[2] The establishment of such a polarity during compaction is thought to generate an environmental identity for inside and outside cells of the embryo. Consequently, stochastic expression of the above transcription factors is amplified into a feedback loop that specifies outside cells to a TE fate and inside cells to an ICM fate. In the model, an apical environment turns on Cdx2, which upregulates its own expression through a downstream transcription factor, Elf5. In concert with a third transcription factor, Eomes, these genes act to suppress pluripotency genes like Oct4 and Nanog in the outside cells.[2][8] Thus, TE becomes specified and differentiates. Inside cells, however, do not turn on the Cdx2 gene, and express high levels of Oct4, Nanog, and Sox2,[2][3] These genes suppress Cdx2 and the inside cells maintain pluripotency generate the ICM and eventually the rest of the embryo proper.
Although this dichotomy of genetic interactions is clearly required to divide the blastomeres of the mouse embryo into both the ICM and TE identities, the initiation of these feedback loops remains under debate. Whether they are established stochastically or through an even earlier asymmetry is unclear, and current research seeks to identify earlier markers of asymmetry. For example, some research correlates the first two cleavages during embryogenesis with respect to the prospective animal and vegetal poles with ultimate specification. The asymmetric division of epigenetic information during these first two cleavages, and the orientation and order in which they occur, may contribute to a cell’s position either inside or outside the morula,.[11][12]
Stem cells
Blastomeres isolated from the ICM of mammalian embryos and grown in culture are known as embryonic stem (ES) cells. These pluripotent cells, when grown in a carefully coordinated media, can give rise to all three germ layers (ectoderm, endoderm, and mesoderm) of the adult body.[13] For example, the transcription factor LIF4 is required for mouse ES cells to be maintained in vitro.[14] Blastomeres are dissociated from an isolated ICM in an early blastocyst, and their transcriptional code governed by Oct4, Sox2, and Nanog helps maintain an undifferentiated state.
One benefit to the regulative nature in which mammalian embryos develop is the manipulation of blastomeres of the ICM to generate knockout mice. In mouse, mutations in a gene of interest can be introduced retrovirally into cultured ES cells, and these can be reintroduced into the ICM of an intact embryo. The result is a chimeric mouse, which develops with a portion of its cells containing the ES cell genome. The aim of such a procedure is to incorporate the mutated gene into the germ line of the mouse such that its progeny will be missing one or both alleles of the gene of interest. Geneticists widely take advantage of this ICM manipulation technique in studying the function of genes in the mammalian system,.[1][13]
Additional images
 Blastodermic vesicle of Vespertilio murinus.
Blastodermic vesicle of Vespertilio murinus. Section through embryonic disk of Vespertilio murinus.
Section through embryonic disk of Vespertilio murinus.
See also
References
- 1 2 3 Wolpert, Lewis. Principles of Development: Third Edition. 2007. Oxford University Press.
- 1 2 3 4 5 6 Marikawa, Yusuke, et al. Establishment of Trophectoderm and Inner Cell Mass Lineages in the Mouse Embryo. Molecular Reproduction & Development 76:1019–1032 (2009)
- 1 2 Suwinska A, Czołowska R, Ozdze_nski W, Tarkowski AK. 2008. Blastomeres of the mouse embryo lose totipotency after the fifth cleavage division: Expression of Cdx2 and Oct4 and developmental potential of inner and outer blastomeres of 16- and 32-cell embryos. Dev Biol 322:133–144.
- ↑ Koyama et al Analysis of Polarity of Bovine and Rabbit Embryos by Scanning Electron Microscopy Biol of Reproduction, 50, 163-170 1994
- ↑ Kuijk, et al Validation of reference genes for quantitative RT-PCR studies in porcine oocytes and preimplantation embryos BMC Developmental Biology 2007, 7:58 doi:10.1186/1471-213X-7-58
- 1 2 Nichols J, Zevnik B, Anastassiadis K, Niwa H, Klewe-Nebenius D, Chambers I, Sch€oler H, Smith A. 1998. Formation of pluripotent stem cells in the mammalian embryo depends on the POU transcription factor Oct4. Cell 95:379–391.
- ↑ Rodda DJ, Chew JL, Lim LH, Loh YH, Wang B, Ng HH, Robson P. 2005. Transcriptional regulation of nanog by OCT4 and SOX2. J Biol Chem 280:24731–24737.
- 1 2 Strumpf D, Mao CA, Yamanaka Y, Ralston A, Chawengsaksophak K, Beck F, Rossant J. 2005. Cdx2 is required for correct cell fate specification and differentiation of trophectoderm in the mouse blastocyst. Development 132:2093–2102.
- ↑ Nishioka N, Yamamoto S, Kiyonari H, Sato H, Sawada A, Ota M, Nakao K, Sasaki H. 2008. Tead4 is required for specification of trophectoderm in pre-implantation mouse embryos. Mech Dev 125:270–283.
- ↑ Nishioka N, et al. 2009. The Hippo signaling pathway components Lats and Yap pattern Tead4 activity to distinguish mouse trophectoderm from inner cell mass. Dev Cell 16: 398–410.
- ↑ Bischoff, Marcus, et al. Formation of the embryonic-abembryonic axis of the mouse blastocyst: relationships between orientation of early cleavage divisions and pattern of symmetric/asymmetric divisions. Development 135, 953-962 (2008)
- ↑ Jedrusik, Agnieszka, et al. Role of Cdx2 and cell polarity in cell allocation and specification of trophectoderm and inner cell mass in the mouse embryo. Genes Dev. 2008 22: 2692-2706
- 1 2 Robertson, Elizabeth , et al. Germ-line transmission of genes introduced into cultured pluripotential cells by retroviral vector. Nature 323, 445 - 448 (2 October 1986)
- ↑ Smith AG, Heath JK, Donaldson DD, Wong GG, Moreau J, Stahl M and Rogers D (1988) Inhibition of pluripotential embryonic stem cell differentiation by purified polypeptides. Nature, 336, 688–690
External links
- Thomas A. Marino, Ph.D. - Embryology Lectures, Temple university (archive)
- Week 1: Implantation